father
Input Quality Assessment
father.1_phred33
father.2_phred33
Statistics
| source: father_result/father_bwamem.sort.rmdup.readfiltered.bam | |
| QC-passed reads | 81093904 |
| QC-failed-reads | 0 |
| Duplicates | 4395852 |
| Mapped reads | 72469565(89.36%) |
| Paired reads | 81093904 |
| Read1 | 40546952 |
| Read2 | 40546952 |
| Properly paired | 79384298(97.89%) |
| Reads with itself and mate mapped | 72443713 |
| Singletons | 25852(0.03%) |
| Reads with mate mapped to a different chromosome | 470431 |
| Reads with mate mapped to a different chr (mapQ at least 5) | 470431 |
| source: father_result/father_bwamem.sort.rmdup.readfiltered.bam | |
| total length (defined by /home/yunfeiguo/projects/seqmule_parallel/bin/secondary/../../misc/hg19_exome.bed) | 46.21Mb |
| Fraction of reads mapped to target region | 53.22% |
| Average coverage in target region | 69.91 |
| Percentage above 30 | 73.03% |
| Percentage above 20 | 81.25% |
| Percentage above 10 | 89.25% |
| Percentage above 5 | 93.29% |
| source: father_result/father.extract_consensus.vcf | |
| Sample | father |
| Number of variants | 31184 |
| Number of SNPs | 29494 |
| Number of indels | 1690 |
| Transitions | 21860 |
| Transversions | 7650 |
| Ti/Tv Ratio | 2.86 |
| Total heterozygotes | 18553 |
| Ref/Alt heterozygotes | 18509 |
| Alt/Alt heterozygotes | 44 |
| Homozygotes | 12631 |
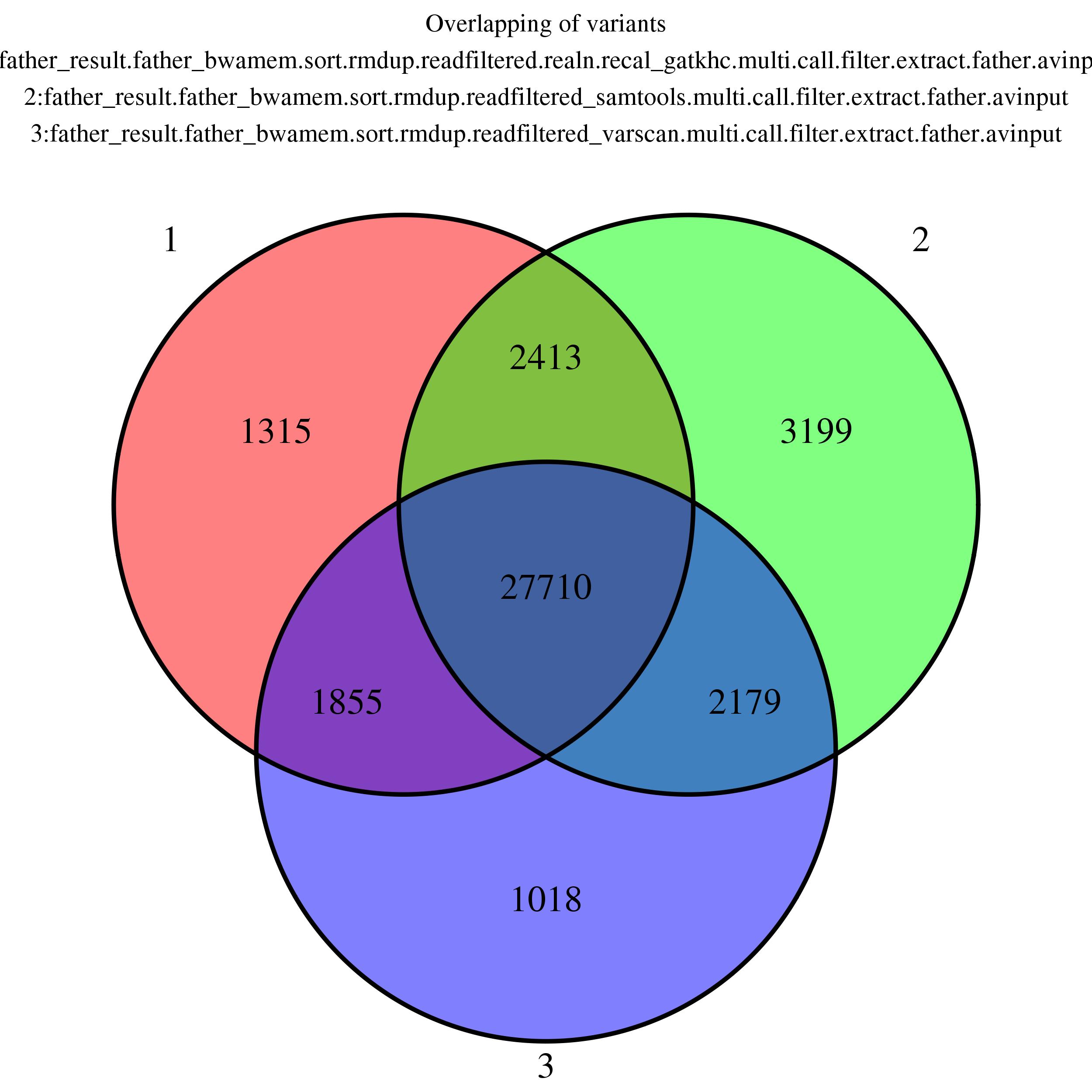
Venn Diagram
Showing overlapping of up to 5 variant output
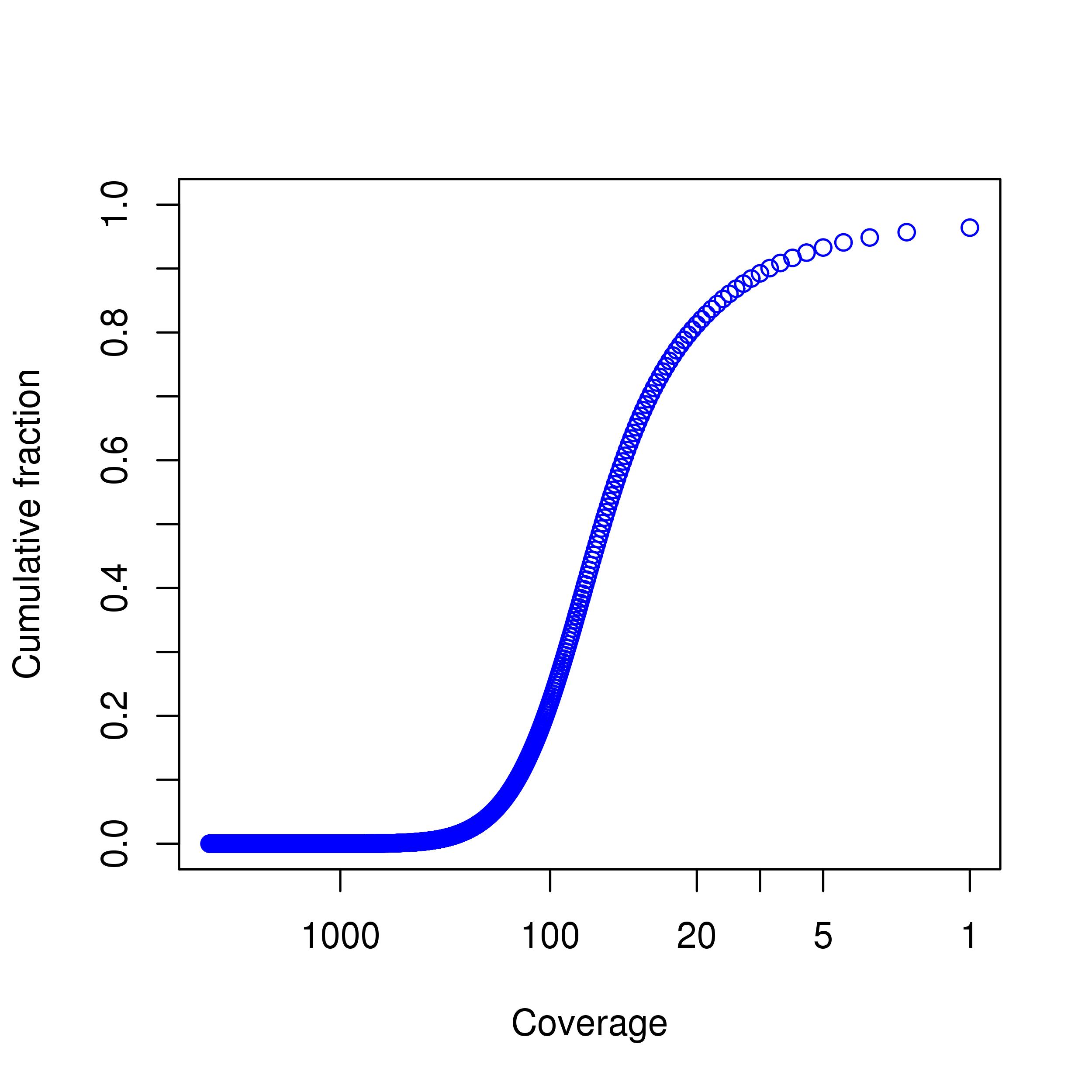
Coverage Plot