Install PennCNV main package
Below is the instruction for installing PennCNV on various operating systems. Although pre-compiled executables are included in the software package, they may not match the specific system architecture that users may have; therefore, re-compilation of the source code is always recommended. In fact, many of the problems that users reported can be simply solved by re-compilation of the source code!
1 Installation on various operating systems
1.1 Linux system
1.2 Installation under Windows - overview
1.3 Windows system with Cygwin
1.4 Windows system with ActivePerl
1.5 Mac OS X
1.6 Compilation from source
2 Installation trouble shooting
2.1 Problem (out of memory in Cygwin):
2.2 Problem (program is very slow):
2.3 Problem (wave adjustment does not work):
2.4 Problem (“core dumped” or “cannot locate loadable object” error):
2.5 Problem (cannot find -lperl):
2.6 Problem (PennCNV does not compile on Ubuntu):
Installation on various operating systems
If using Linux/Unix system, we can run the wget command to download the program files in a local directory:
[kai@adenine tutorial1 ENS40]$ wget http://www.openbioinformatics.org/penncnv/download/penncnv.latest.tar.gz
Then we uncompress the downloaded file by the tar command, which automatically generate the penncnv/ directory containing executables and library files:
[kai@adenine tutorial1 ENS40]$ tar xvfz penncnv.latest.tar.gz
We will see the new directory called penncnv/, enter this directory, and we will see several Perl programs (file name in the *.pl format), as well as a kext/, lib/ and example/ subdirectory. Next, cd into the kext/ directory, optionally make chagne to the Makefile, and type “make”. If there is no error message, the installation is done!
Now try to run the detect_cnv.pl program:
[kai@adenine penncnv ENS40]$ ./detect_cnv.pl
It will show the program usage information, indicating the successful installation of the program. If an error message like “/usr/bin/perl: bad interpreter: No such file or directory” is shown, it indicates that Perl is installed in a different directory: try running “perl ./detect_cnv.pl” instead. If other types of error message are issued (such as floating point exception), we need to compile the program, see the “compilation” section below.
If your computer is not a standard PC with a standard installation of Perl/Linux, to ensure that PennCNV runs correctly in your system, always try to re-compile the source code! See the more detailed instruction below.
For Solaris, change "gcc" to "cc" in the Makefile, since normally one can create shared libraries with c compiler provided by Solaris, but not gcc itself.
ADDITIONAL NOTES: If you have problem installing PennCNV in your operating system, it is perhaps due to incompatibility with the latest version of Perl 5.14/5.18 in some systems. To solve this issue, you can use perlbrew to install a lower version of Perl and run PennCNV on top of that without re-compilation. Perlbrew defaults the installation to non-threaded and non-multi perl versions. In order to use (for example), as precompiled code for perl-5.8.8 (x86_64-linux-thread-multi, one would have to set their perlbrew install to install a version where thread and multi is available. So, in the install I had to do "perlbrew install perl-[insert version here] --thread --multi". See more instructions at the end of this page.
Installation under Windows - overview
There are two ways to use PennCNV in a Windows system: either install Cygwin and use PennCNV under Cygwin shell, or install ActivePerl and use PennCNV in a regular Windows command shell. These two methods are described in the following two sections.
If you really like to use Windows, and if you do not want to use command line, you can alternatively try to use PennCNV inside BeadStudio software by the Universal CNV adapter. You can access the webpage describing this procedure in the PennCNV Plug-in page.
In this section we describe the method for running PennCNV under Cygwin.
We first need to download and save the PennCNV package as zip file from a web browser to local hard drive, and use a unzipping software to unzip the files. Do make sure the kext/, lib/ and example/ directories are present after unzipping, since some software under Windows is unable to generate directory structures from *.tar.gz file.
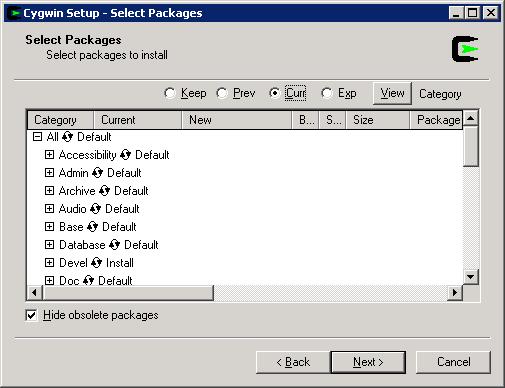
We next need to install cygwin in the system. Do make sure that you select the “devel” package during cygwin installation, since the “devel” package is NOT selected in default installation process. (Basically by clicking the word next to “devel” in the package-selection dialog, you will see the option cycle through “default”, “reinstall”, “uninstall” and “install”: stop clicking when seeing “install” as the option and then go the next installation step (see figure below). Depending on your network connection speed and computer speed, it may take 5-30 minutes for downloading the package and another 5-10 minutes to install the package.
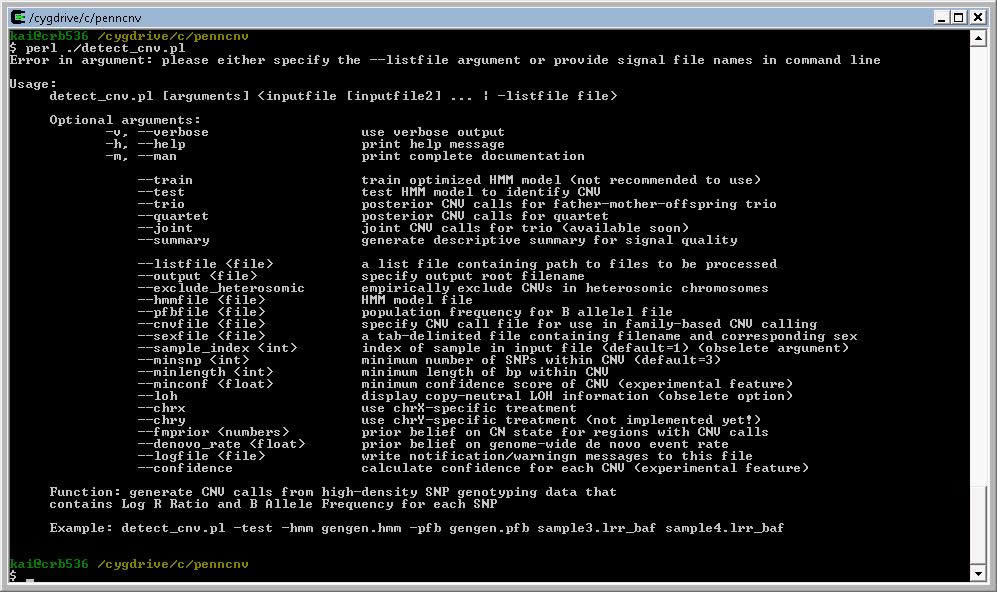
After cygwin is installed in your system, you can then open the Cygwin BASH shell and run PennCNV program from there. Note that in cygwin, to access the penncnv/ directory in local hard drive, we need to use a pathname similar to “cd /cygdrive/c/penncnv”, rather than “cd c:\penncnv”. When running the detect_cnv.pl program, if you see an error message, it possibly indicates that the devel package has not been installed in cygwin environment.
Windows system with ActivePerl
Note: Current PennCNV package contains pre-compiled executables for 32-bit Perl in Windows XP and Windows Vista and Windows 7. If you have a 64-bit Windows system, you should install a 32-bit Perl ( 5.8.9, 5.10.1, 5.12.3, 5.14.0) in the 64-bit Windows, and PennCNV should be able to run in compatibility mode.
Like the last section, we first need to download and uncompress the PennCNV package. We next need to install ActivePerl in the computer: go to http://www.activeperl.com, click a few buttons to download the free version of ActivePerl . Install the file to your Windows system by selecting all default options. Previous versions of PennCNV only supports Perl 5.8, but the new version now supports more Perl versions.
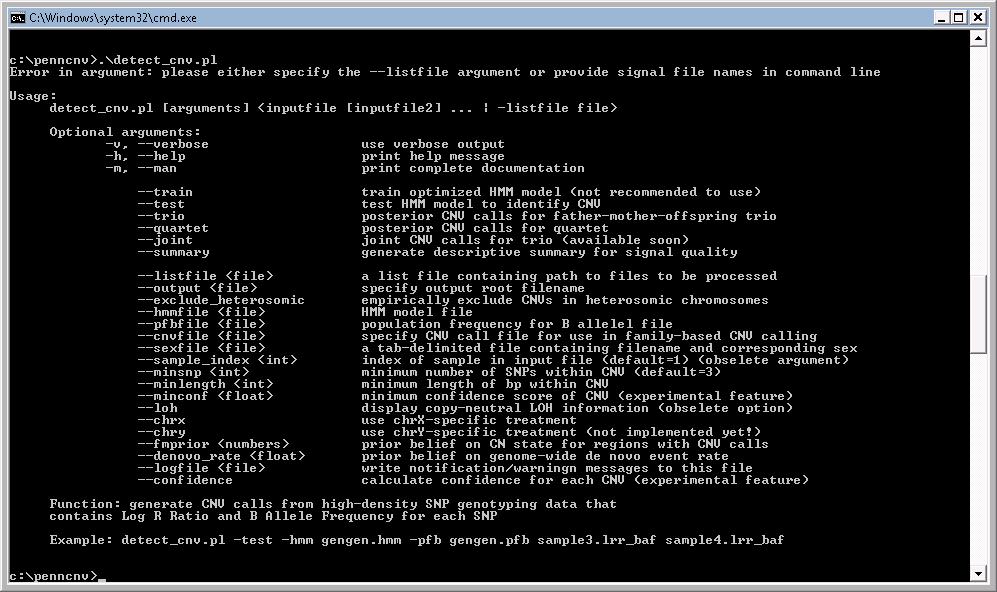
Now we can open a Windows command shell (click the “start”, then click “Run…”, then type in “cmd”, then type enter). Go to the directory where PennCNV is installed (for example, “cd c:\penncnv”), then type “perl .\detect_cnv.pl”, and we should see the program usage information here. In fact, we can also try typing “.\detect_cnv.pl” directly, and it should also work (if ActivePerl is installed with default options for handling *.pl extensions). See example below.
Users need to recompile the source code for execution in Mac OS. This information was provided by Markus Ringner: In the Makefile, replace “-shared” with “-dynamiclib”, replace “khmm.so” with “khmm.dylib”.
If you see a “core dumped” error message, or see a “cannot locate loadable object” error message, when running detect_cnv.pl program, it means that the executable is not compatible with your system architecture. If using Linux system, please download the source code and re-compile executable by these steps: (1) unzip the source tar.gz file (2) cd into the kext/ directory (3) optionally edit the “Makefile” to change options (4) type “make” (5) If there is no error message, the installation is done!
If using a very old machine with very old version of “make” program, the fancy characters like $@ and $^ in the Makefile may not be recognized correctly. If that’s the case, just manually change the character to appropriate file names.
Compilation in Cygwin: the procedure is slightly different: change khmm.so to khmm.dll in the Makefile file before compilation.
Compilation in Windows: The pre-compiled executable in PennCNV can only be used under 32-bit Windows machine. (If you use 64-bit Windows, it is likely that the pre-compiled executable can still be used, but only if you install 32-bit Perl in the machine, not the 64-bit version of Perl). If you have a 64-bit Windows but have no experience in compilation, do NOT compile. Just download 32-bit Perl and run PennCNV directly. I was not able to compile PennCNV specifically for 64-bit Perl.
For other Windows system and/or any other Perl version, re-compilation may be required. Follow these steps exactly. First, install ActivePerl in the system (direct link to 32-bit version of 5.8.9, 5.10.1, 5.12.3, 5.14.0), and it does not matter which version you choose to install. Next, install the MinGW-Get (minimalist GNU), which contains necessary components/commands/libraries for GCC. During installation, select the MSYS shell which is more convenient for our compilation purposes, compared to windows shell. Next, open MSYS, go to PennCNV directory, to go the kext sub-directory, run the perl -MExtUtils::Embed -e ccopts, perl -MExtUtils::Embed -e ldopts , perl -MConfig -e 'print $Config{archname}' and perl -MConfig -e 'print $Config{version}' command, then copy the output and paste into the Makefile in the corresponding places in the file (make sure to delete the ` sign). Different machines may have a different outputs for the above commands! (for example, some machines will need to use VER=5.8.8 but others will need to use VER=5.10.1, depending on version of Perl). The only complication is that any “\” character need to be changed to “/” instead. For example, “C:\Perl\lib\CORE” should be changed to “C:/Perl/lib/CORE”. Finally change khmm.so to khmm.dll in the Makefile file. Now just type “make” in the kext/ directory and it should work from there.
Compilation in Solaris: change "gcc" to "cc" in the "Makefile", then compile by typing "make". This change is needed because gcc in many Solaris system is not compiled in a way to allow dynamic linking.
Problem (out of memory in Cygwin):
Symptom: “out of memory” occurs in cygwin, although the machine clearly has 4GB memory and should be more than enough (even for 1.8 million markers for a father-mother-offspring trio, the memory usage should be ~3GB in a 64-bit machine).
Solution: By default, Cygwin assigns only 1GB memory so any process, so a process that tries use >1GB will be killed by Cygwin.
Try this in cygwin:
regtool -i set /HKLM/Software/Cygnus\ Solutions/Cygwin/heap_chunk_in_mb 2048
regtool -v list /HKLM/Software/Cygnus\ Solutions/Cygwin
If it does not work, change the 2048 to 4096. If this still does not work, then I do not know a good solution, except to split the PFB file into two parts (one for chr1-10, the other for 11-23), and run PennCNV twice with two different PFB files.
Problem (program is very slow):
Symptom: Program takes forever to run.
Solution: It should take ~2 minutes for calling CNV on a signal file with 1 million markers, and slightly longer if the GCmodel adjustment and confidence score calculation are used. If the program takes forever to run, it is more likely due to lack of memory. In Windows, use the task manager to check the CPU/memory usage; in Linux/Unix, use the “top” command to check CPU/memory usage. If the CPU usage is far less than 20%, it indicates that virtual memory has been used by the system to execute PennCNV, resulting in the slow speed. To solve the problem, try to use a machine with larger amounts of memory instead. Alternatively, split the PFB file into two parts (one for chr1-10, the other for 11-23), and run PennCNV twice with two different PFB files.
Problem (wave adjustment does not work):
Symptom: The -gcmodel argument does not work, so the GCWF score before and after adjustment are the same.
Solution: Re-compile the source code using the instruction above.
Problem (“core dumped” or “cannot locate loadable object” error):
Symptom: When issuing PennCNV, the “cannot locate loadable object” message appear, or when running PennCNV, a sudden “core dumped” message occurs during what appears to be a normal run.
Solution: Re-compile the source code using the instruction above.
Symptom: when typing "make" to compile the program, gcc complains that " /usr/bin/ld: cannot find -lperl"
Solution: The error is caused by the fact that perl is not installed in a
standard way so the path to libperl is not annotated in the "perl -MExtUtils::Embed -e ldopts" command output.
If you run "perl -V",
you can try to find something like"/usr/lib64/perl5/5.8.8/x86_64-linux-thread-multi/CORE" in the dynamic
linking section. Now make sure that the"/usr/lib64/perl5/5.8.8/x86_64-linux-thread-multi/CORE/libperl.so"
file actually exist. Then in the Makefile, just add"-L/usr/lib64/perl5/5.8.8/x86_64-linux-thread-multi/CORE/"
to the command it should fix the problem.
To give a more clear example, suppose after using "perl -V", you
know that the library "libperl.so" file is in the "/usr/lib" folder.
But if you run "perl -MExtUtils::Embed -e ldopts" you found the
following: "
-Wl,-E -L/usr/local/lib -L/usr/lib/perl/5.10/CORE -lperl -ldl -lm
-lpthread -lc -lcrypt", so /usr/lib is not annotated here.
Now you can just simply manually add the "-L/usr/bin" to the command line, "
gcc -shared -o khmm.so khmm_wrap.o khmm.o kc.o include/arrays.o khmmDev.o -L/usr/lib -Wl,-E -L/usr/local/lib -L/usr/lib/perl/5.10/CORE
-lperl -ldl -lm -lpthread -lc -lcrypt",
so the system won't complain about not finding "-lperl".
Problem (PennCNV does not compile on Ubuntu)
Symptom: compilation of PennCNV gives error message in Ubuntu
Solution: The following instruction is given by Dan Porat: If you use Ubuntu, note that the default installation of Ubuntu does not contain libperl-dev and therefore it should be added. The only step needed in order to get PennCNV up and running is the command:"sudo apt-get install libperl-dev".
If you still cannot make it work in Ubuntu, then follow these suggestions: you can use perlbrew to install a lower version of Perl and run PennCNV on top of that without re-compilation. Perlbrew defaults the installation to non-threaded and non-multi perl versions. In order to use (for example), as precompiled code for perl-5.8.8 (x86_64-linux-thread-multi, one would have to set their perlbrew install to install a version where thread and multi is available. So, in the install I had to do "perlbrew install perl-[insert version here] --thread --multi".
# If installing pennCNV with the standard procedure does not work
# (http://www.openbioinformatics.org/penncnv/penncnv_installation.html)
# eg on Ubuntu 14.04 with perl 5.18.2, architecture x86_64-linux-gnu-thread-multi:
# This might be due to incompatibilities of pennCNV's khmm module with recent perl versions
# Solution: install an older perl with the perlbrew utility (which will not erase your current perl)
# and use precompiled khmm for perl 5.8.8 available in the pennCNV download
## INSTALL compatible version of perl with the perlbrew utility
# install perlbrew
\wget -O - http://install.perlbrew.pl | bash
source ~/perl5/perlbrew/etc/bashrc
# install perl 5.8.8
perlbrew install perl-5.8.8 --thread --multi # says that installation failed because 1 test failed
cd /home/yourhomename/perl5/perlbrew/build/perl-5.8.8; make install # force install anyway
## USAGE: run detect_cnv.pl
# first possibility requiring perlbrew:
source ~/perl5/perlbrew/etc/bashrc # loads perlbrew
perlbrew use perl-5.8.8 # assigns perl to perl-5.8.8
perl /path/to/pennCNV/detect_cnv.pl --test --hmm example.hmm --pfb example.pfb father.txt mother.txt offspring.txt # run pennCNV
# Note:
# the following command will not work even if the pennCNV folder is in your $PATH,
# because the #!/usr/bin/perl at the beginning of detect_cnv.pl points to the system perl and not to perl-5.8.8
# (even if you ran the perlbrew use perl-5.8.8 command)
detect_cnv.pl --test --hmm example.hmm --pfb example.pfb father.txt mother.txt offspring.txt
# second possibility: forget about perlbrew:
# replace the first line of detect_cnv.pl : #!/usr/bin/perl
# by the location where perlbrew installed perl-5.8.8 : #!/home/yourhomename/perl5/perlbrew/perls/perl-5.8.8/bin/perl
# then the following command works (if the pennCNV folder is in your $PATH):
detect_cnv.pl --test --hmm example.hmm --pfb example.pfb father.txt mother.txt offspring.txt